Identifying genes that regulate the cellular arrangement and composition of a tumor microenvironment (TME) through which cancers facilitate their unchecked growth is a vital goal of researchers. Drugs that target components of the TME, such as immune checkpoint blockade and angiogenesis inhibitors, can be very effective anticancer therapies.
Still, identifying genetic determinants of the TME has been a herculean task because it requires perturbing one gene at a time and learning how that impacts the cellular architecture and immunosuppressive characteristics of the tumor in vivo. These gene-by-gene studies are informative, but not very efficient given the urgent need for new cancer therapies.
A next-generation spatial genomic technique developed by a TCI team could represent a quantum leap in that discovery process. The technology, known as Perturb-map, allows researchers to knock out dozens, or even hundreds, of genes in parallel and simultaneously assess through imaging how each knockout influences tumor growth, histopathology, immune composition, and molecular state.

From left to right, Ashwitha Lakshmi, BSc, MSc; Miriam Merad, MD, PhD; Brian Brown, PhD; and Jaime Mateus-Tique, BSc, MSc.
“The idea of scaling up from one gene at a time to 50 or 100 genes gives us the ability to study and identify, in a way we never could before, specific genes that control the extracellular environment of the tumor and the immune cells surrounding it,” says Brian Brown, PhD, a TCI member, Director of the Icahn Genomics Institute and Associate Director of the Marc and Jennifer Lipschultz Precision Immunology Institute at Mount Sinai. The technique and its findings were published in Cell in March 2022, with Dr. Brown as senior author.
“Perturb-map essentially allows us to expose the tumor’s vulnerabilities, and that is the key to helping us find new targets for cancer therapy, particularly immunotherapy.”
Brian Brown, PhD
Scientists are accustomed to using CRISPR, the gene editing tool, to knock out genes in cancer cells for large-scale studies. Perturb-map builds on that technique through a novel system that essentially “paints” cells with different color barcodes and then identifies cancerous cells with gene modifications, as well as neighboring noncancerous cells within the tumor.
In their investigation, TCI researchers used these CRISPR genetic screens to knock out 35 genes in parallel in a mouse model of lung cancer. Among the questions they sought to answer was how tumors orchestrate their characteristic patterns, shapes, and local cell environments.
“Cancer cells protect themselves from the immune system by turning genes off or on, which controls whether T cells penetrate the tumors and are able to kill the malignant cells,” explains Dr. Brown. “We wanted to learn which specific genes they are because they represent potential targets of vulnerability for the right drugs. That’s where Perturb-map is so valuable: It allows us to identify those genes.”
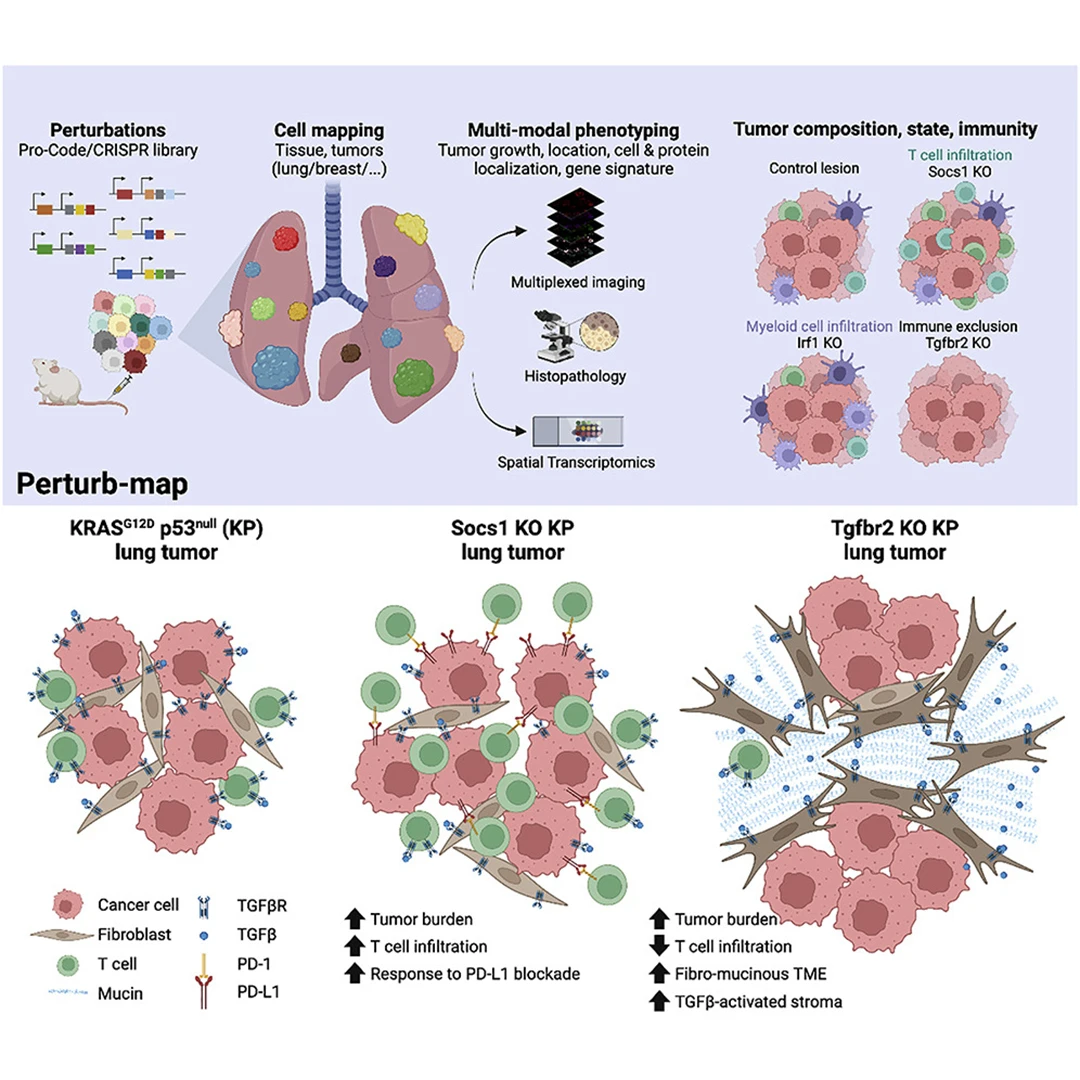
An illustration showing how Perturb-map's color coding mapping works (top), and how in a Tgfbr2 knockout tumor model, functional genomics can be performed within the tissue at single-cell resolution with spatial architecture preserved, with insights into the tumor microenviroment (bottom).
Specifically, Perturb-map enabled the team to discover several key pathways that have major effects on tumor growth and architecture, as well as immune cell recruitment. One pathway is controlled by the cytokine interferon gamma (IFNγ), and another by the tumor growth factor beta (TGFβ) receptor. What they found was when the SOCS1 gene, which codes for suppressor of cytokine signaling 1 (SOCS1) protein, a negative regulator of IFNγ, was deleted from cancer cells, the lung tumors became larger and more abundant.
Furthermore, the research revealed that SOCS1 gene knockout tumors were highly infiltrated by T cells, while the TGFβ receptor gene knockout tumors excluded T cells—an important finding since patients whose tumors contain fewer T cells respond more poorly to immunotherapy drugs.
These findings also underscored for Dr. Brown that the effects of these genes on the tumor ecosystem are extremely localized. “This is a notable insight because we’re learning that many patient tumors are composed of genetically distinct subclones,” he says.

The work on Perturb-map from Dr. Brown's and Dr. Merad's teams can knock out dozens, or even hundreds, of genes in parallel, whereas current methods have to do them one by one.
If specific gene mutations are keeping T cells out of a subclonal region, it might serve as a pocket of resistance to current immunotherapies such as Keytruda and Tecentriq, which turn on T cells in the tumor so they can kill cancer cells. Thus, if a tumor region has no T cells, it means the drugs have no cells to turn on, and the tumor will not be eliminated. Based on that rationale, the study suggested that SOCS1 might be a good therapeutic target to pair with PD1/PD-L1 blockade, since SOCS1 inhibition could enhance T cell infiltration while blunting the effects of upregulated PD-L1.
Dr. Brown emphasizes the long-range potential of functional genomic technologies such as Perturb-map. “By revealing how cancer cells control the extracellular environment of the tumor at scale,” he says, “the technique greatly accelerates our ability to find new therapeutic targets—and that is key to improving cancer outcomes.”
Featured

Brian Brown, PhD
Director, Icahn Genomics Institute; Associate Director of PrIISM

Miriam Merad, MD, PhD
Mount Sinai Professor in Cancer Immunology; Director of PrIISM