Looking at the results of an RNA sequence analysis of early-stage lung adenocarcinoma (esLUAD) samples, Charles A. Powell, MD, MBA, and a team of Mount Sinai researchers noticed an interesting phenomenon. The algorithm separated the tumors based on gene expression into two groups—one distinguished by genes associated with aggressive tumors (e.g., acinar, micropapillary, papillary, and solid) and one distinguished by genes that are associated with indolent tumors.
Was it possible, they wondered, that these gene signatures could predict outcomes among individuals with early-stage lung cancer tumors that looked the same on chest CT scan?
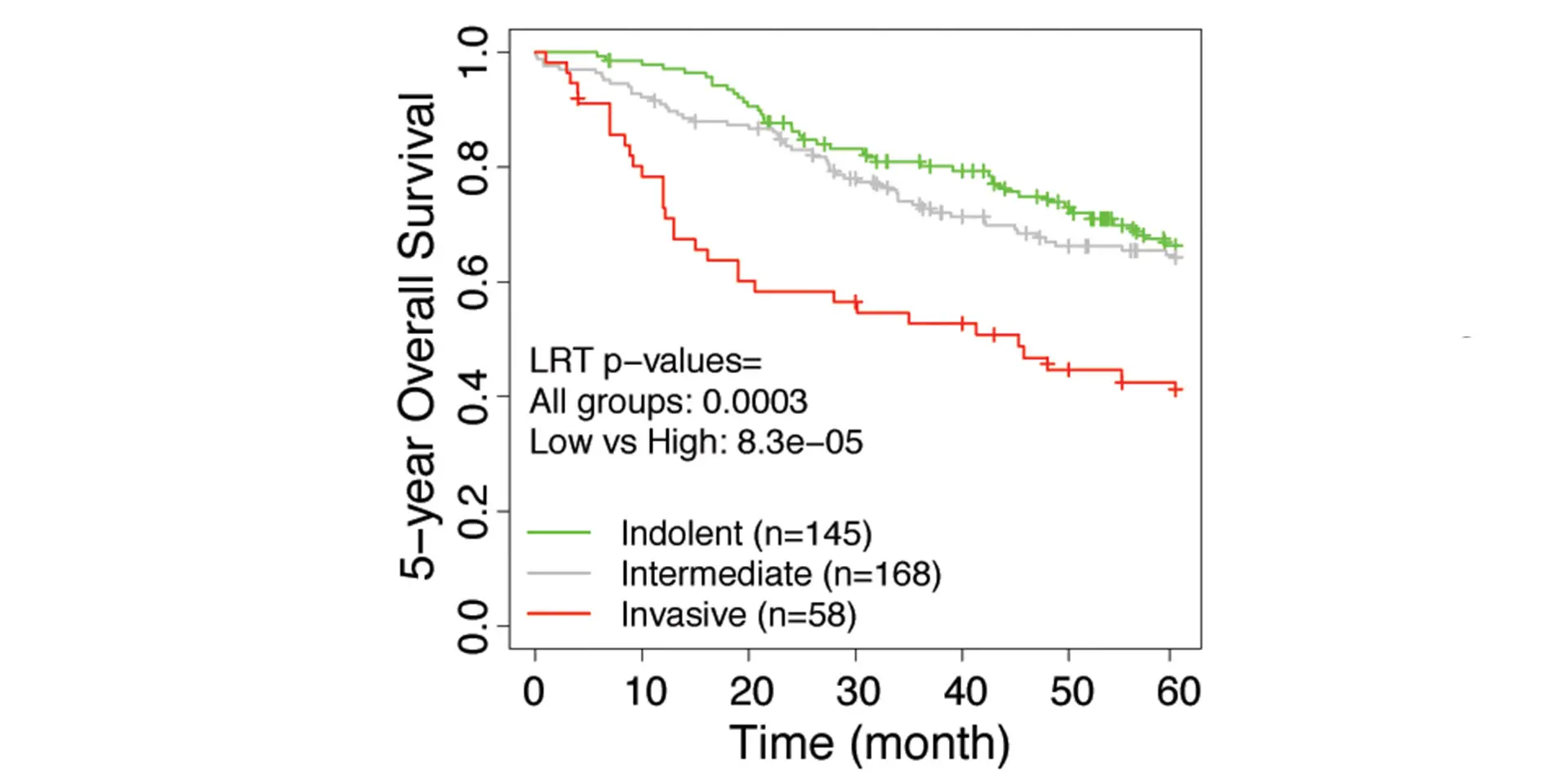
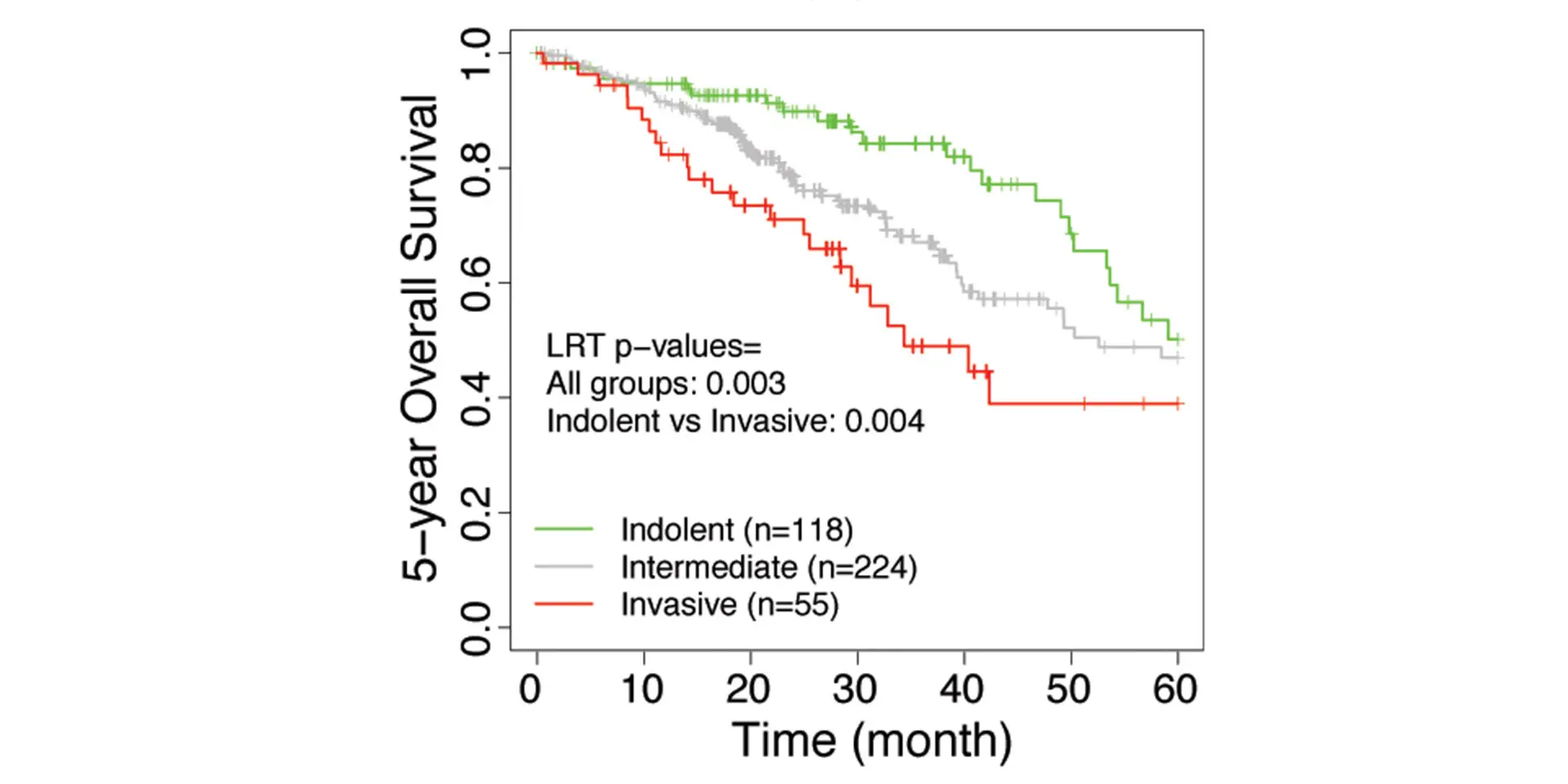
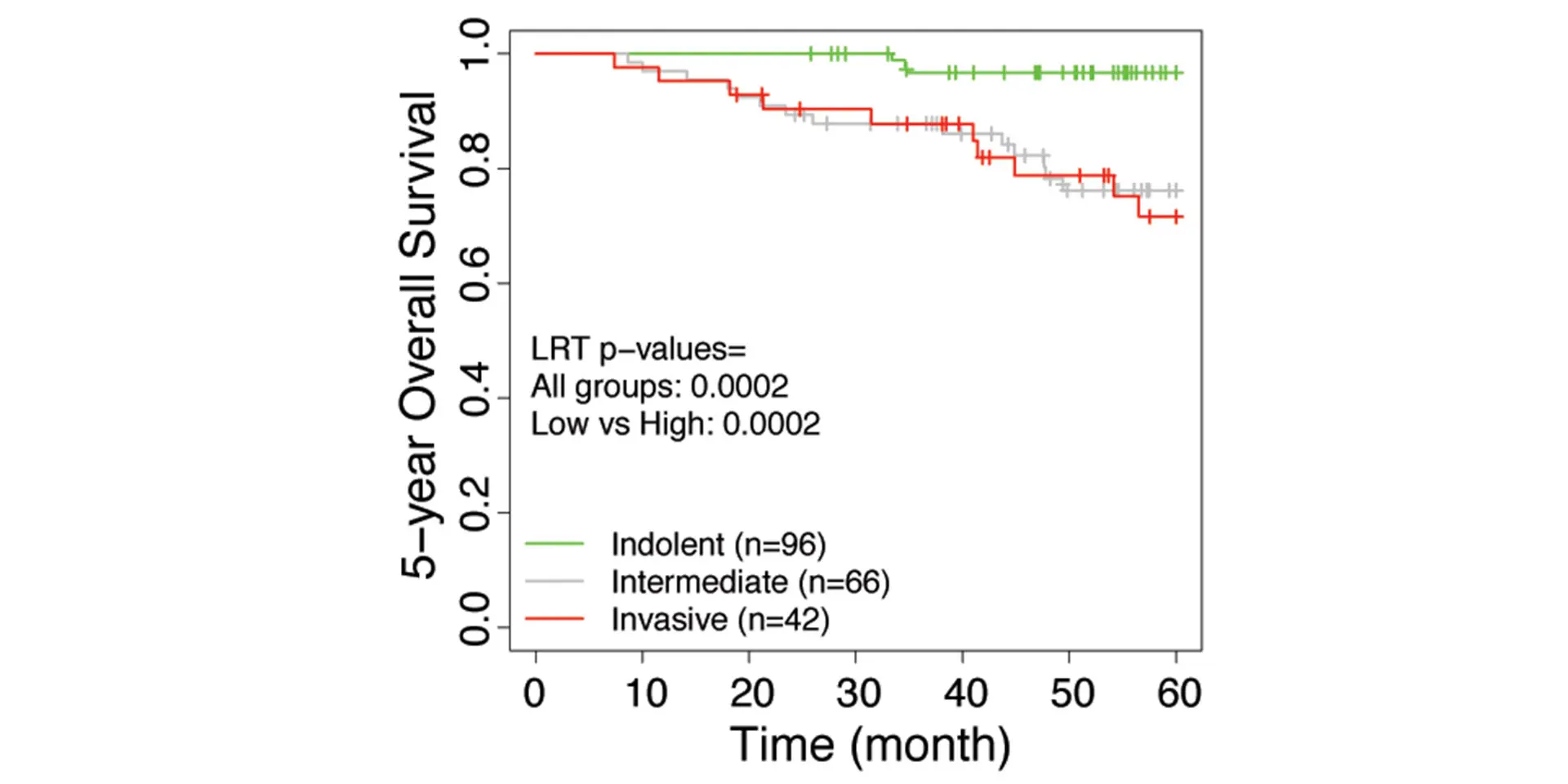
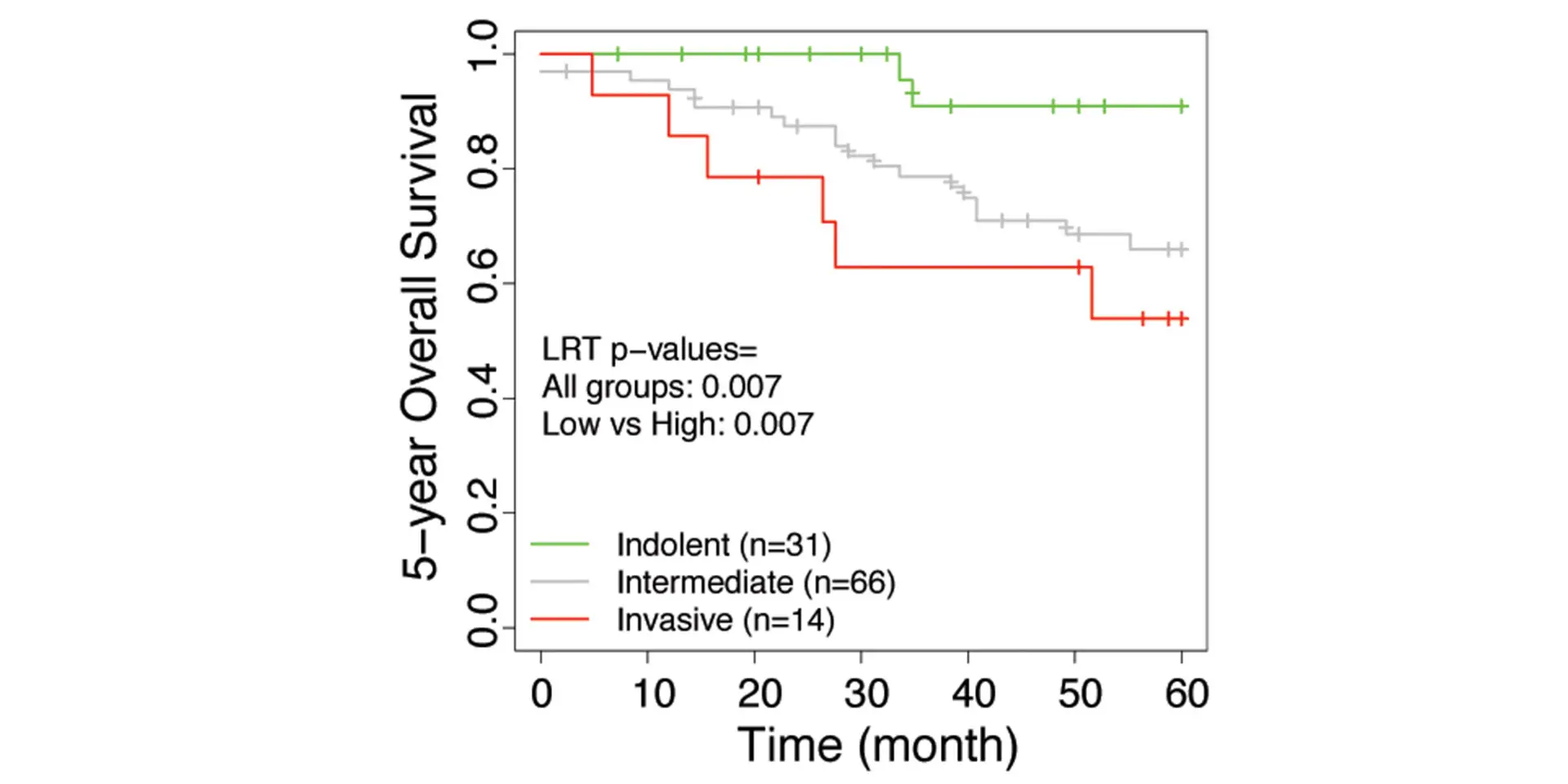
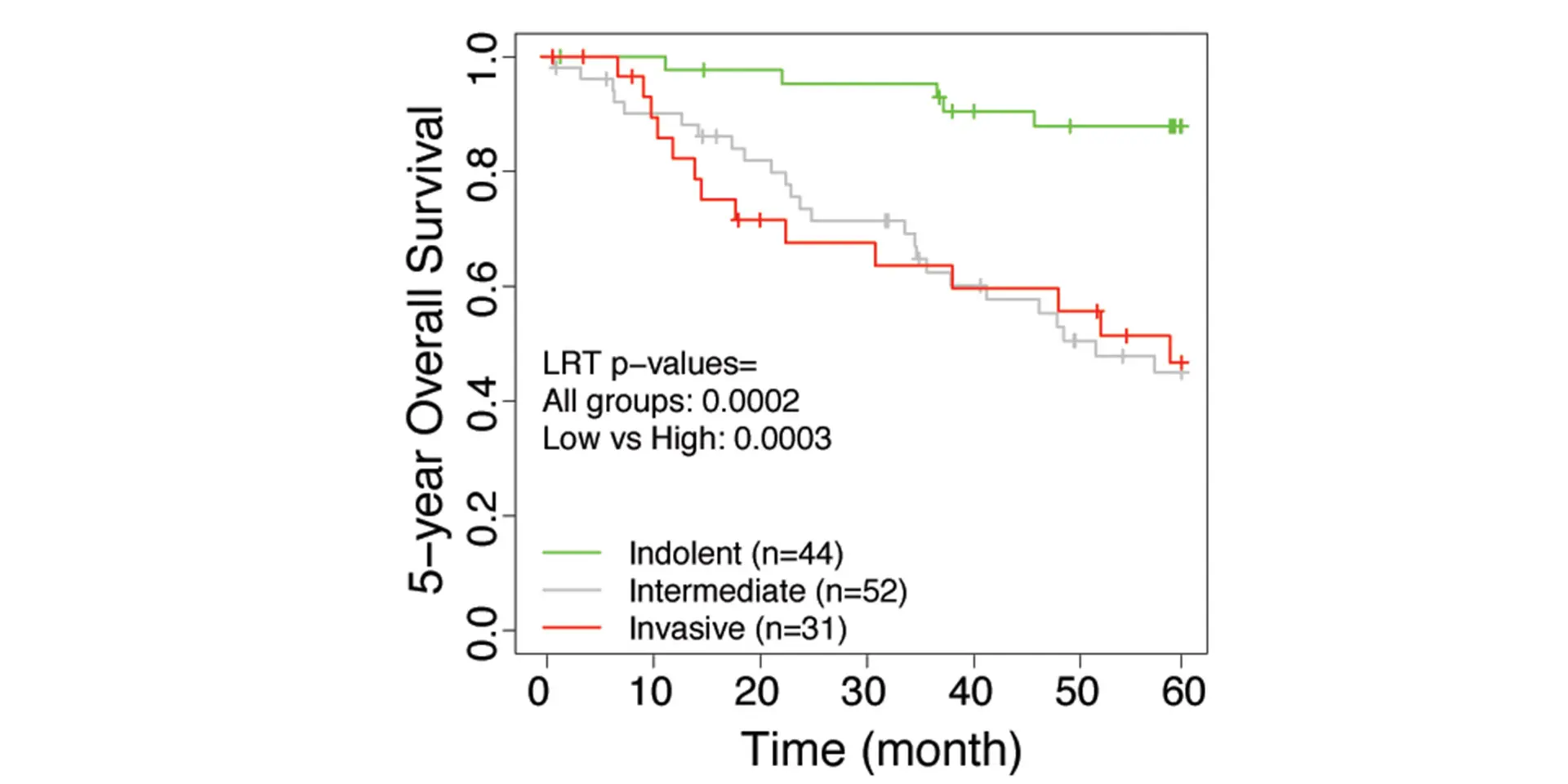
The research team developed an invasiveness score (IVS) and applied the signatures to seven published data sets from the United States, Europe, and Japan that have included the gene expression of lung adenocarcinoma patients with their associated survival rates. Based on the IVS, samples were classified as indolent, invasive, or intermediate. For all seven data sets, the researchers found that the invasive groups had significantly inferior survival compared to the indolent groups. Furthermore, the IVS remained significantly associated with survival in these cohorts after adjustment for age, gender, and tumor stages. The study, “Integrative Network Analysis of Early-Stage Lung Adenocarcinoma Identifies Aurora Kinase Inhibition as Interceptor of Invasion and Progression,” was published March 24, 2022, in Nature Communications.
“This indicates that there is the potential for gene signatures derived from lung carcinoma to predict outcomes among individuals with small lung cancers detected by chest CT imaging,” says Dr. Powell, Director, Mount Sinai – National Jewish Health Respiratory Institute, and Janice and Coleman Rabin Professor of Pulmonary Medicine at the Icahn School of Medicine at Mount Sinai.
A New Way to Assess a Tumor's Indolence
For years, imaging and histology have been critical considerations in the management of patients with esLUAD. If the computed tomography (CT) scan and the microscope sample had the appearance of an aggressive tumor, it was treated with medication or surgery. If the tumor appeared indolent, it was managed through a watchful waiting approach. But Dr. Powell knows that appearances can be deceiving.
“Tumors that have a predicted indolent course on the basis of imaging may harbor genetic and genomic alterations that suggest an increased risk for those tumors to progress to and cause an adverse outcome,” Dr. Powell says. “We thought that if we combine an analysis of the molecular features of these tumors with their imaging characteristics, we could better refine the probability of these lesions actually being and remaining indolent versus the probability of them changing their behavior and necessitating a different management strategy.”
To test that theory, Dr. Powell and his team acquired and conducted molecular profiling of a cohort of 53 resected esLUAD specimens with histology classification that included adenocarcinoma in situ and minimally invasive adenocarcinoma—a criterion that distinguished the data set from other larger data sets that do not have that volume of indolent tumors available for analysis.
“We had two hypotheses coming into this project,” Dr. Powell says. “One was that the genomic signatures of these tumors would distinguish themselves into distinct subsets that would correlate with the extent of invasion detected pathologically in these tumors. The second hypothesis was that we would be able to identify key drivers of the networks that were responsible for the molecular and biological changes that would distinguish these tumors.”
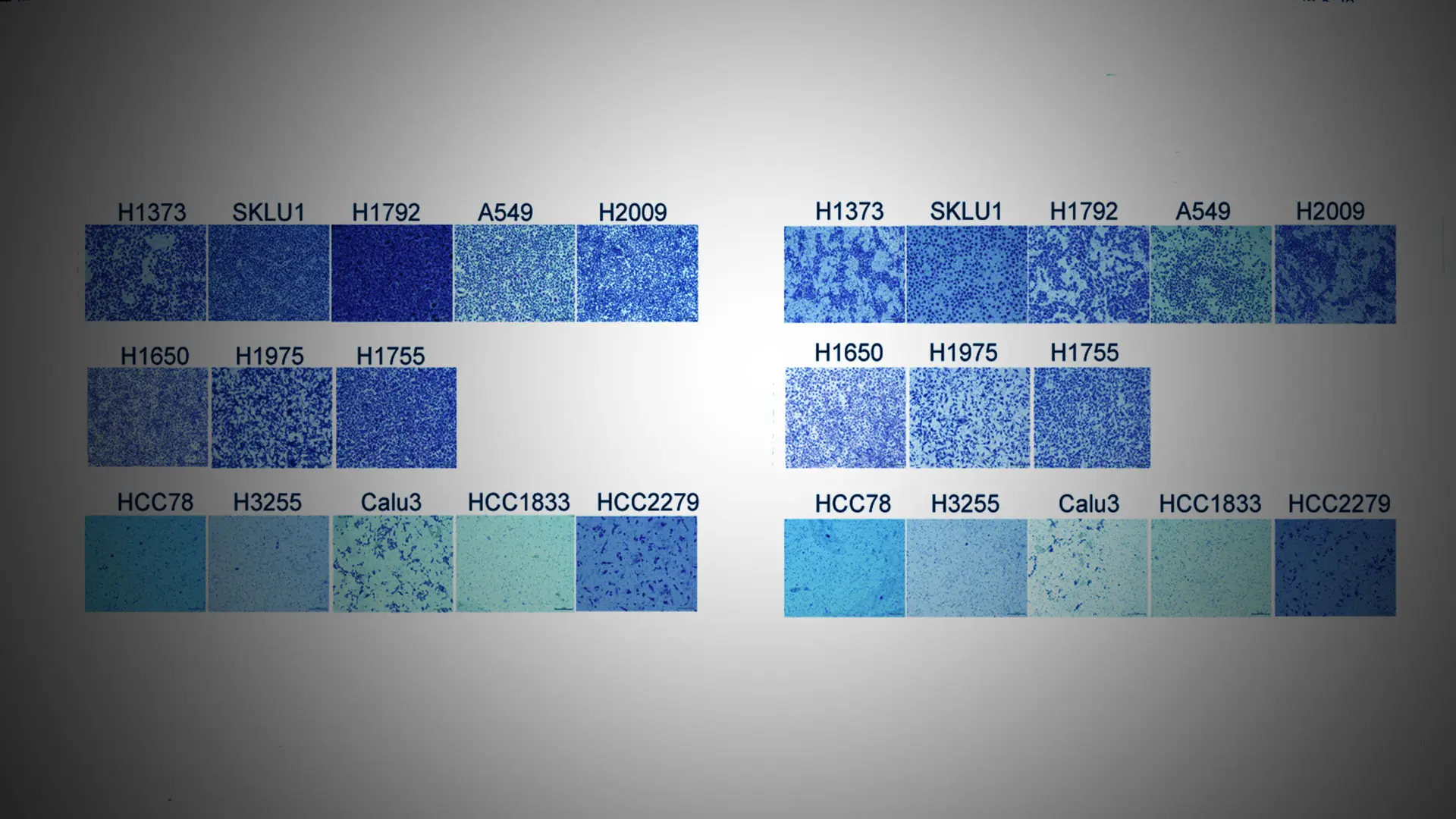
Having validated the first hypothesis through collaboration with the Department of Genetics and Genomic Sciences at Icahn Mount Sinai and Sema4, a Mount Sinai spin-out, Dr. Powell and the research team undertook a network analysis of in vitro cancer cell systems to explore how the 1,322 differentially expressed genes they identified were connected, identify the gene signatures that were most important in cancer progression, and identify key drivers of the genetic signatures. They identified the aurora kinase A (AURKA) and aurora kinase B (AURKB) proteins as the key regulators of the pro-invasive gene signature. The team then evaluated the behavior of the cancer cells in terms of their invasion potential by eliminating both aurora kinase signaling pathways through gene deletion and small molecule inhibitors, including AMG900, which is in clinical trials for use in treating acute myeloid leukemia, advanced solid tumors, and triple negative breast cancer in combination with angiogenic kinase inhibitor ENMD2076. Using both methods, the researchers were able to restore the invasive cells to non-invasive behavior. This network analysis was part of the Nature Communications paper.
“Previously, researchers looked at AURKA and AURKB mainly from the perspective of proliferation,” says Jun Zhu, PhD, Professor, Genetics and Genomic Sciences, at Icahn Mount Sinai. “Because our study was focused on invasiveness, we showed that you need to knock out both proteins to have an impact, which AMG900 is able to do.”
The researchers then investigated the preclinical utility and suppression efficacy of AMG900 using mouse models with transgenic invasive lung adenocarcinoma. In that model, the researchers activated the Kirsten rat sarcoma virus (KRAS) gene—a key driver in 33 percent of lung adenocarcinoma cases—and removed expression of the transforming growth factor, beta receptor II (TGFBR2), resulting in a model of lung adenocarcinoma that progresses from indolent to aggressive. Micro-CT images demonstrated clear differences in tumor density and burden in the treatment group versus the control group, with the control group experiencing a significantly faster increase in tumor burden. Histopathological analysis identified several invasive and in situ tumors with bulky nodules. Although solid and acinar patterns increased among the control animal lungs, the lungs of the treated mice showed suppression of invasiveness. These results were also part of the Nature Communications paper.
“Studies show that tumor metastases occur at an early stage,” Dr. Zhu says. “Our findings are important because by identifying these drivers and demonstrating that inhibition retards the progression of invasion, they open the door for potential early treatment of metastases.”
Dr. Powell cautions that more studies are necessary to assess the significance of these findings. He and his lab are collaborating with Sema4 on a clinical trial that will explore the prognostic and therapeutic potential of the identified signatures and drivers. “We are looking at administering aurora kinase inhibitors among patients with suspected esLUAD in a neoadjuvant approach to assess the impact in terms of gene signatures and tumor pathology,” he says.
Featured

Charles A. Powell, MD, MBA
Chief of Pulmonary, Critical Care and Sleep Medicine; Director, Mount Sinai Respiratory Institute

Jun Zhu, PhD
Professor of Genetics and Genomic Sciences