Highlighting the contributions made to the field by Panagiotis (Panos) Roussos, MD, PhD; Towfique Raj, PhD; Ian S. Maze, PhD; and Schahram Akbarian, MD, PhD
In 2005, the emergence of second- (or next-) generation sequencing marked a significant advance in genomic research, enabling the integration of large-scale genome sequencing and surveillance into a wide range of experimental applications.
By facilitating the capture of genomic variation associated with both common and rare diseases, next-generation sequencing has become a powerful tool for studying the underlying genetic basis of a wide range of neurological and psychiatric disorders. It has also markedly promoted the field’s study of the molecular basis of brain function, such as transcriptomes and epigenomes, by allowing their profiling in a comprehensive and unbiased manner. This has greatly facilitated our understanding of the regulation of gene expression, and the impact of environmental factors on gene function, across the life cycle.
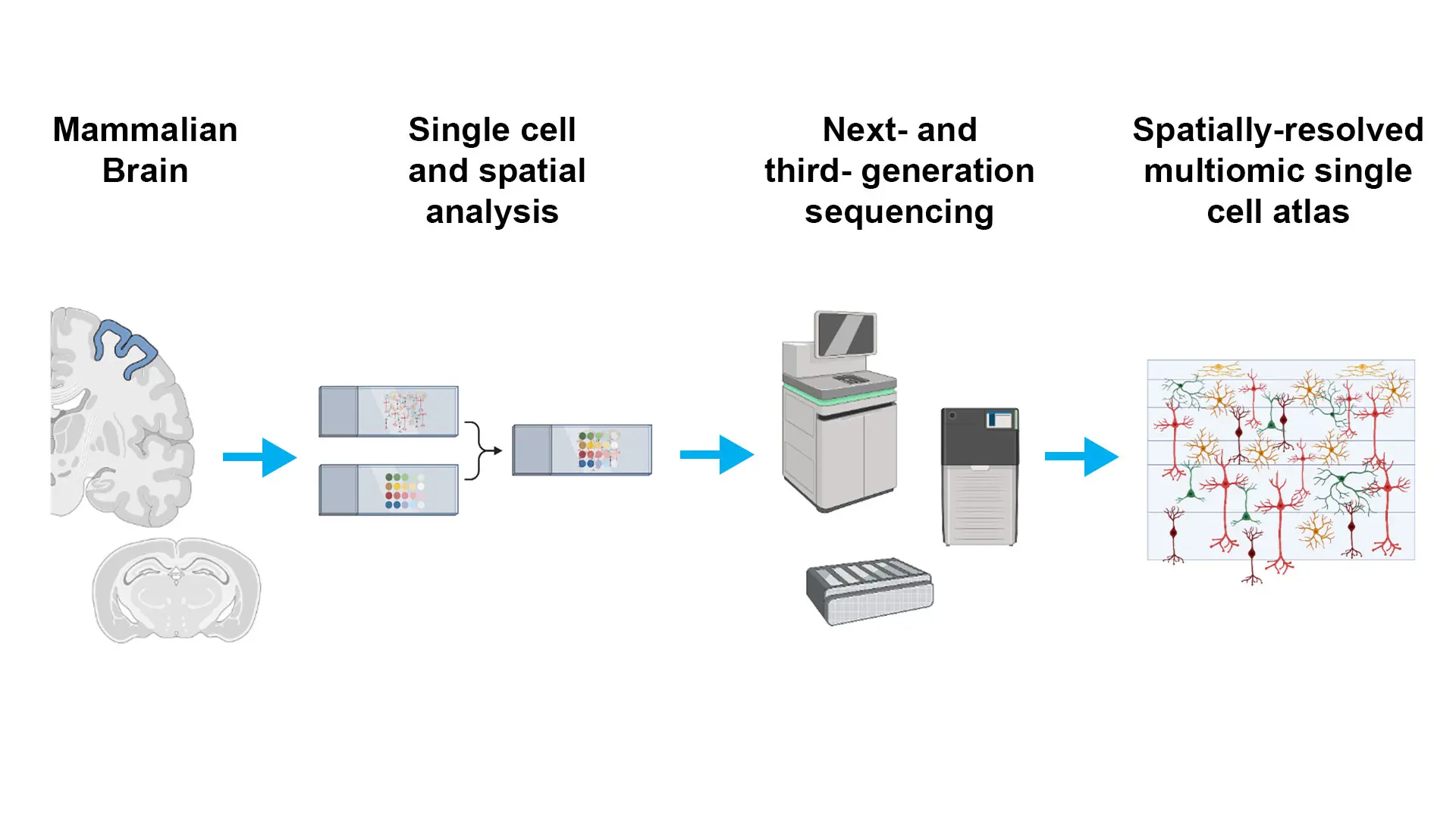
Figure 1. The combination of next-generation sequencing and single-cell analyses is revolutionizing our ability to perform multiomic profiling with single-cell spatial resolution, providing an unprecedented level of insight into the complex biology of the mammalian (especially human) brain. These technologies hold enormous promise for advancing our understanding of human health and disease, and they are likely to continue to be a driving force in neuroscience research for years to come.
In the early 2010s, third-generation (or long) sequencing was introduced, enabling the direct sequencing of entire (many kilobase-long) nucleic acid molecules. This innovative technology generates significantly longer reads than its next-generation counterpart, making it possible to detect DNA and RNA modifications on nucleic acids with much higher accuracy. Overall, the development of next- and third-generation sequencing technologies has had a transformative effect on the field, opening up new avenues for the study of genetic and epigenetic mechanisms underlying normal development of the human brain and abnormalities throughout life that underlie central nervous system disorders (see Figure 1).
Over the last decade, advances in single-cell sequencing technologies—encompassing DNA, RNA, and epigenomics—have represented paramount technological breakthroughs. The complexity of the mammalian brain, with its enormous number of neurons and glia and the unanticipated diversity seen for individual cell types, presents a unique challenge for researchers aiming to understand the underlying mechanisms of neural circuit function. Accurately characterizing these cell types, as well as their roles in functionally distinct regions of the brain, is crucial for developing a comprehensive understanding of brain function (see Figure 1).
Prior to the advent of single-cell sequencing technologies, most genomic, transcriptomic, and epigenomic measurements reported an averaged signal that blended multiple cell types from brain tissues. This made it challenging to unravel cellular heterogeneity and comprehend the regulatory mechanisms of gene expression programs in distinct subpopulations of cells. The emergence of single-cell sequencing has transformed the ways in which we profile tissues by providing an unbiased, marker-free approach to distinguish and analyze the myriad cell types present in the brain and the pattern of gene expression they exhibit.
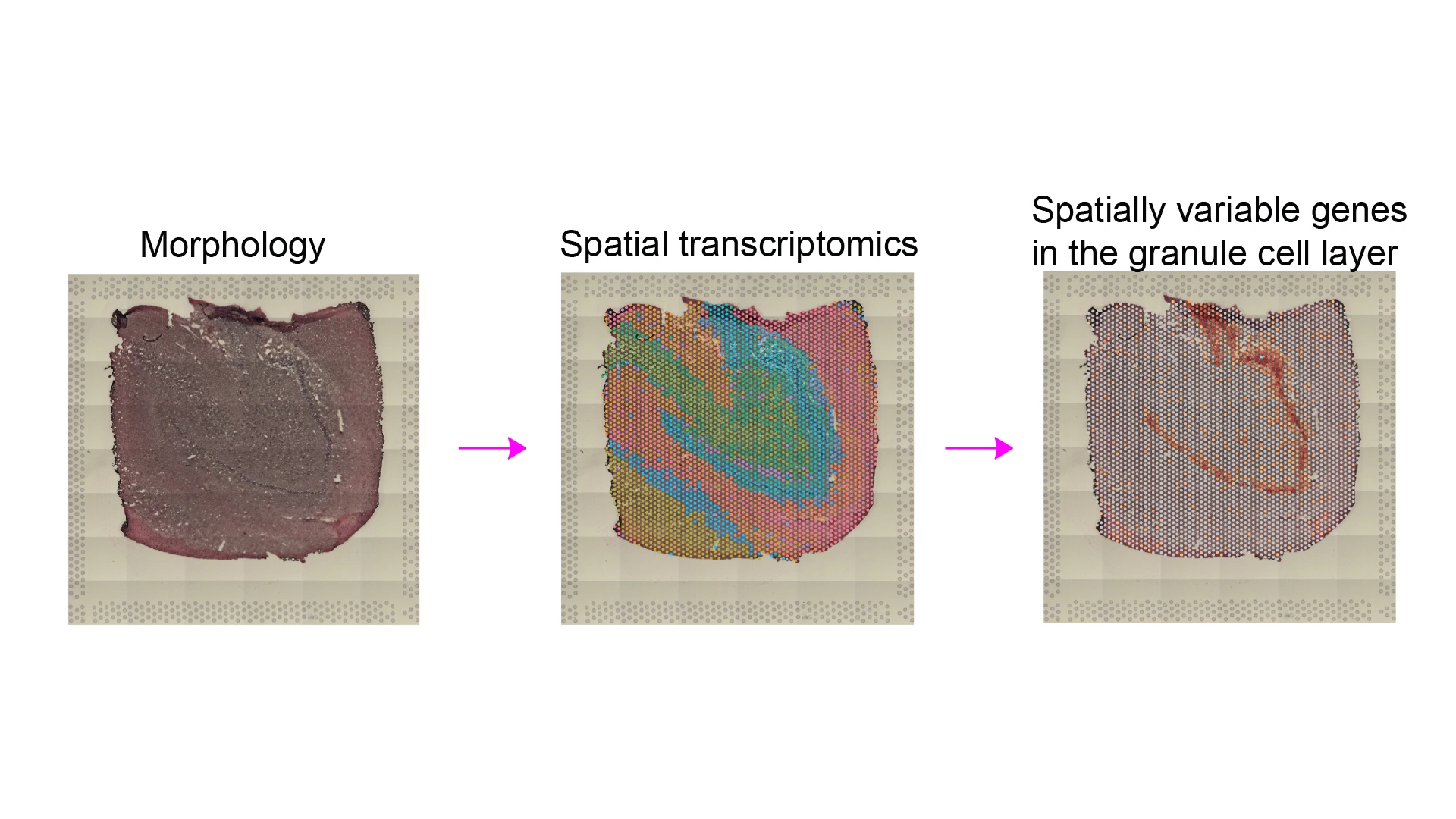
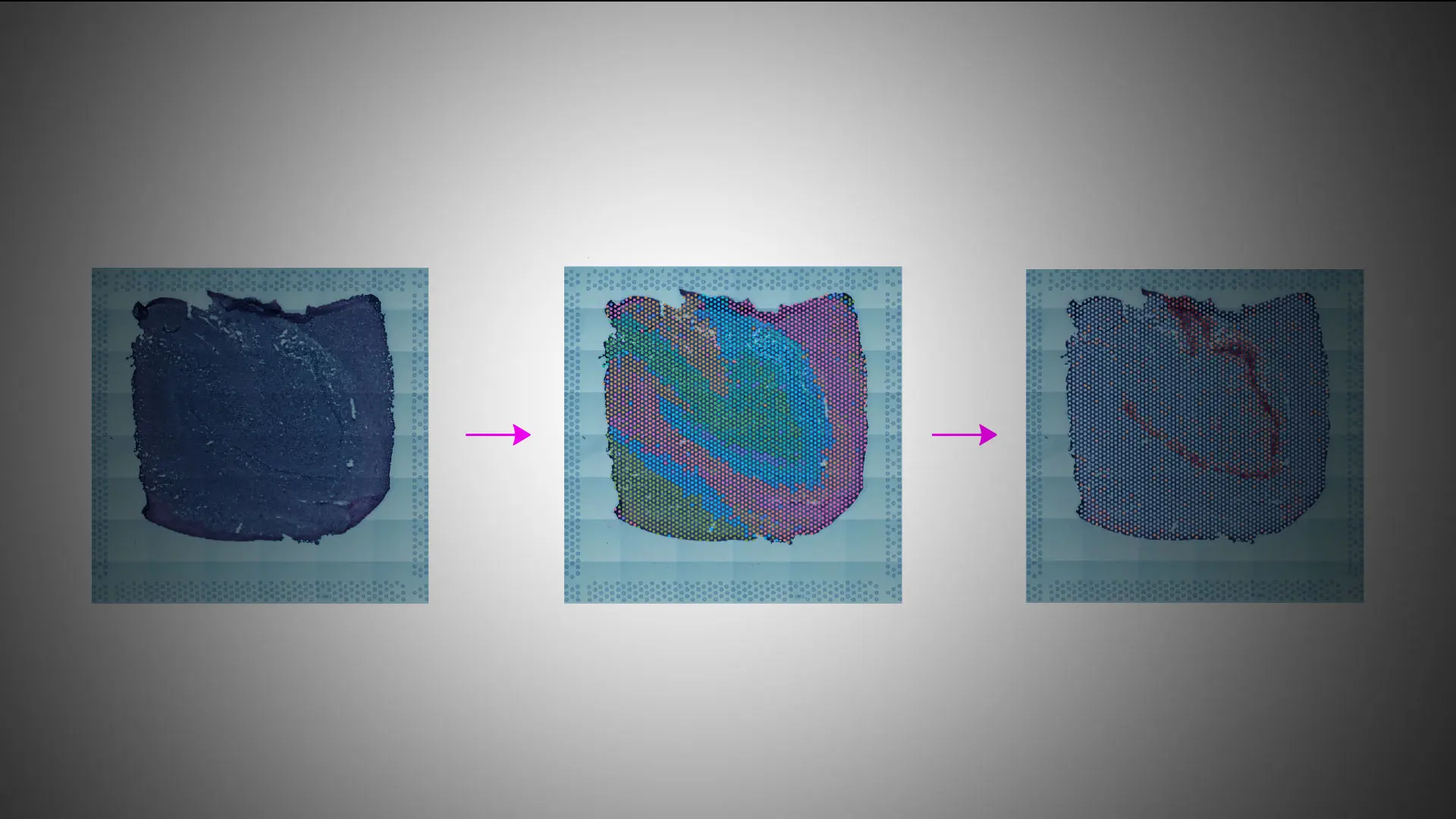
Figure 2. Morphology (H&E staining), spatial transcriptomics using 10x Genomics Visium platform, and spatially variable genes detected in the granule cell layer of the human dentate gyrus.
Spatial genomics, the ability to determine gene expression patterns within a given cell type in the context of its location in intact tissue, has emerged as a further technological advance in more recent years. This technology allows researchers to study cells as they normally exist within tissues and organs and provides important insights into cell-cell interactions, their spatial organization, and the roles of distinct cell types within the functional context of a given tissue (see Figure 2).
These various platforms are now being applied to many hundreds of brains from people with a range of neurological or psychiatric disorders and from associated animal models. In the next 10 years, it is expected that further transformative advances in neuroscience will be made as these methods achieve low-cost, high-throughput multiomic profiling at single-cell spatial resolution and are applied to an even larger number of human and animal samples.
Such a platform will provide new insights into genotype-phenotype interactions and link the fields of histopathology and imaging to single-cell genomics, opening novel avenues of investigation in diverse fields of biology and disease. These advances will greatly improve our understanding of the complexity of the brain and enable the development of more targeted therapies for a wide range of common and rare neurodevelopmental and neurodegenerative disorders.
Featured

Panagiotis (Panos) Roussos, MD, PhD
Professor of Genetics and Genomic Sciences, and Psychiatry, and Director of the Center for Disease Neurogenomics

Towfique Raj, PhD
Associate Professor of Neuroscience, Genetics and Genomic Sciences, and Neurology

Ian S. Maze, PhD
Professor of Neuroscience, and Pharmacological Sciences, and Director of the Center for Neural Epigenome Engineering

Schahram Akbarian, MD, PhD
Professor of Genetics and Genomic Sciences, Neuroscience, and Psychiatry, and Chief of the Division of Psychiatric Epigenomics
